Gallery

Cover Art. Structural studies of Mcm10, a component of the vertebrate replisome

Cover Art. TIBS review on new DNA repair mechanisms

Cover Art. Discovery of an E. coli system for unhooking interstrand DNA crosslinks
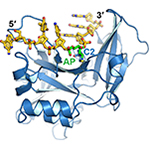
YedK DNA-protein crosslink. Covalent linkage between a SRAP domain and ssDNA containing an abasic site
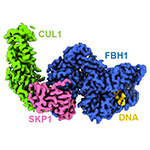
SCFFBH1. Helicase/ubiquitin-ligase complex important for replication fork reversal and RAD51 regulation
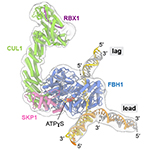
SCFFBH1 bound to fork DNA. FBH1 binds immediately at the fork junction to catalyze replication fork reversal
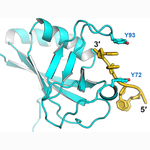
HLTF HIRAN domain. Specific for DNA 3′-ends, the HIRAN domain is critical for replication fork reversal by HLTF
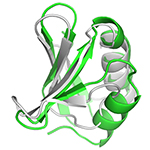
SMARCAL1 HARP domain. Substrate specificity domain of mouse SMARCAL1, involved in repair of stalled replication forks
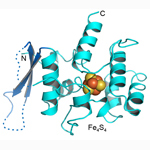
Human pol α–primase PRIM2C iron-sulfur cluster domain. DNA polymerase involved in lagging strand synthesis

Catalytic complexes of DNA pol α–primase. Xenopus pol α–primase in complex with primed templates representing various stages of DNA synthesis.
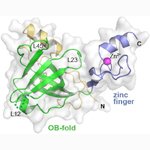
Mcm10 DNA-binding domain. Xenopus laevis Mcm10 OB-fold and Zn-finger domain
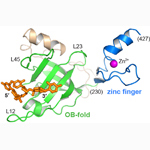
Mcm10-DNA complex. Xenopus laevis Mcm10 internal domain bound to single-stranded DNA
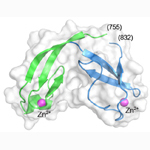
Mcm10 C-terminal domain. NMR structure of the zinc-ribbon domain of Xenopus laevis Mcm10

Mcm10 coiled-coil domain. From Xenopus laevis Mcm10, a protein involved in eukaryotic DNA replication
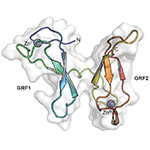
NEIL3 GRF Zinc Finger domain. The tandem GRF-ZF domain of vertebrate NEIL3 DNA glycosylase binds single-stranded DNA
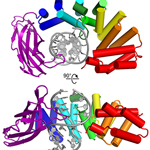
AlkC. HEAT-like repeat 3-methyladenine DNA glycosylase bound to abasic-DNA
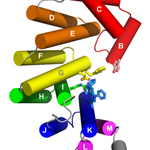
AlkD. Bacillus cerus 3-methyladenine DNA glycosylase (AlkD)
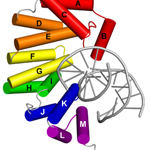
AlkD-DNA complex. Non-catalytic complex of AlkD bound to DNA mismatches
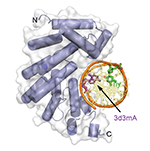
AlkD-DNA catalytic complex.The protein caught in the act of excising 3mA
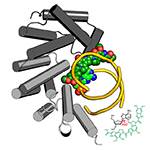
AlkD-Yatakemycin-DNA complex. AlkD excising a DNA adduct of the bulky, genotoxic agent from the duocarmycin/CC-1065 family of natural products
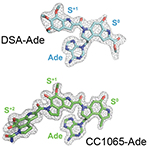
AlkD-Duocarmycin SA-DNA and CC1065-DNA complexes. AlkD excision of Duocarmycin SA and CC-1065 DNA adducts
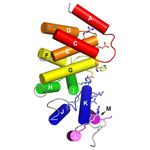
AlkD2. An AlkD homolog from Streptococcus mutans that lacks DNA glycosylase activity
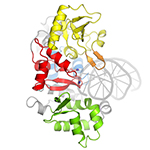
AlkZ. DNA glycosylase that unhooks DNA interstrand crosslinks of the toxin azinomycin B
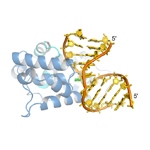
Mag1. S.pombe 3-methyladenine DNA glycosylase bound to damaged DNA
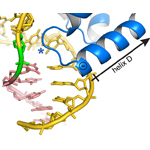
Mag2. Non-catalytic complex of an S.pombe Mag paralog bound to damaged DNA
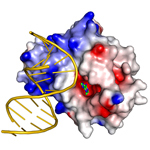
TAG-DNA complex. Salmonella typhi 3-methyladenine (3mA) DNA glycosylase I bound to abasic-DNA and 3mA
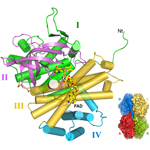
AidB. Non-DNA-repair protein involved in the adaptive response of bacteria to alkylating agents
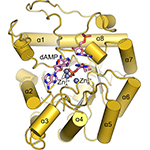
TATDN1. TatD enzymes are evolutionarily conserved AP endonucleases































